BASE - Biologie de l'Adaptation et Systèmes en Évolution
Our research aims at understanding the dynamics of adaptation and the evolution of life-history traits of populations at different biological scales. The associated projects are structured around three main themes: the genotype-phenotype relationship at different levels of phenotypic integration, the dynamics of adaptation at different time scales and interactions between genotype and the biotic and abiotic environment (GxE). To carry out this research, we use experimental approaches (field, greenhouse and laboratory experiments) as well as mathematical modelling approaches. Although most of our projects focus on maize and yeast, we work on other biological models when they are more appropriate for addressing scientific questions, given the technical constraints and the current state of knowledge.
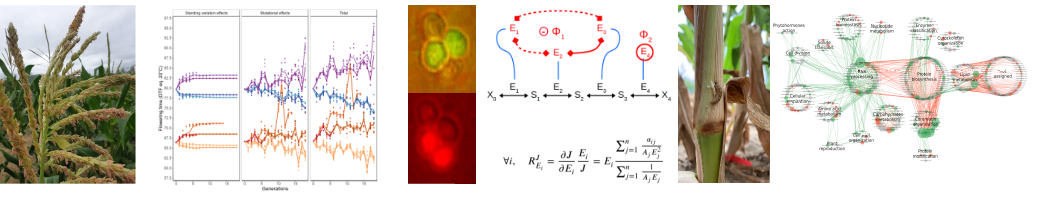
Theme 1. The genotype-phenotype relationship at different levels of phenotypic integration
Understanding the genotype-phenotype (GP) relationship is a central issue in evolutionary biology and plant/animal breeding. The non-linearity of processes shaping this relationship makes it complex.
We are studying this complexity in three projects:
- the basis of heterosis, an emerging property of the complexity of the GP relationship
- the metabolic basis of phenotypic variation in yeast and cultivated plants such as maize
- the evolutionary dynamics of enzyme levels in metabolic pathways. Our approach combines the integration of omics data, mathematical modelling and computer simulations.
Related projects :
- HeteroYeast (ANR), Metaflux (Plant2Pro)
- Current and past PhD projects (date of defense): Maëla Sémery, Yacine Djabali (2023), Charlotte Coton (2021), Marianyela Petrizzelli (2019)
Theme 2. Dynamics of adaptation
o understand the dynamics of adaptation, we study both short- and long-term evolutionary processes. To address these questions, we use both experimental evolutionary and theoretical approaches. This combination of approaches enables us to study how the different scales of life - from molecular mechanisms to populations - interact to shape the dynamics of adaptation.
Our studies mainly deal with :
- the evolution of the rate of recombination and its role in the dynamics of adaptation
- the evolutionary consequences of sexual versus non-sexual reproductive regimes
- the response to selection on flowering time in maize
- survival at a long stationary phase in Escherichia coli as part of a first-year undergraduate teaching unit.
Related projects :
- Evolrec (ANR), CO-PATT (ANR), Itemaize (BASC), EpiFlore (BAP INRAE), Expérience de Sélection Divergente du Plateau de Saclay.
- Current and past PhD projects (date of defense): Arnaud Desbiez-Piat (2021), Elise Tourrette (2020)
Theme 3: Genotype-environment interactions
In the context of global change, a better understanding of the genetic basis of adaptation and GxE interactions is crucial. We are studying the intraspecific genetic and phenotypic diversity of our biological models (plants, micro-organisms) in relation to environmental variations. The response of phenotypes to environmental variations is observed at different scales, from molecular to integrated traits. We are interested in variations in the abiotic environment, in particular drought, and variations in the biotic environment, in particular pest pressure or the rhizospheric microbial communities.
Our studies mainly deal with :
- the diversity of the response of plants (maize, fruit trees) to climatic variables at different life scales
- the adaptation of species to their environments via the identification of genotype-environment associations
- interactions between plants and their biotic environment (insect pests, weeds, pathogenic fungi, microbial communities in the rhizosphere, crop-predator birds, etc.).
Related projects :
- Itemaize (BASC), Amaizing (ANR), WarmRules (DATAIA), INCREASE (Europe), PEPR Agroécologie et Numérique , Stat4Plant (ANR), MYCORE (C-BASC), GAIcirse(C-BASC), PHENOFORE (SEMAE), BIOCOSMA (ANR), MetaP-Rhiz (BAP, Biosphera)
- Current and past PhD projects (date of defense): Maxime Criado, Antoine Caillebotte, Sacha Revillon (2024), Inoussa Sanane (2020)
Members
- Walaa Abdalla PhD student ()
- Diala Abu Awad Assistant professor (Université Paris-Saclay)
- Willy Vincent Bienvenut Junior Investigator (CNRS), 50% BASE, 50% PAPPSO
- Mélisande Blein-Nicolas Research Engineer (INRAE), 50% BASE, 50% PAPPSO
- Monique Bolotin Professor (Université Paris-Saclay)
- Antoine Caillebotte PhD student (INRAE)
- Maxime Criado PhD student (INRAE)
- Dominique de Vienne Professor (Université Paris-Saclay)
- Christine Dillmann Professor (Université Paris-Saclay, Faculté des Sciences)
- Cécile Fairhead Professor (Université Paris-Saclay)
- Matthieu Falque Research Engineer (INRAE), 50% ACEP, 50% BASE
- Nathalie Galic Technician (INRAE), 50% BASE
- Lanah Guin Internship ()
- Marine Le Hingrat Technician (INRAE)
- Judith Legrand Maitre de conférences (Université Paris-Saclay)
- Yulan Liang PhD student ()
- Jévaltonia Lokossou IE, CDD ()
- Élodie Marchadier Assistant Professor (Université Paris-Saclay)
- Thomas Maret Engineer (INRAE)
- Adrienne Ressayre Research scientist (INRAE)
- Maëla Sémery PhD student ()
- You Fang Zhou (Université Paris-Saclay)
Former members
Cédric Bartschi Postdoctoral Fellow (2024-2025), Yasmine Echarafi Research Technician (2023-2024), Xavier Raffoux Engineer (2003-2024), Sacha Revillon PhD student (2021-2024), Yacine Djabali PhD student (2020-2023), Romain Benoist Postdoctoral fellow (2020-2023), Romane Lafontaine Technician (2021-2022), Michel Zivy Directeur de Recherche (1985-2022), Charlotte Coton PhD student (2019-2021), Arnaud Desbiez-Piat PhD student (2018-2021), Adrien Vidal Ingénieur d'études, CDD (2020), Inoussa Sanané PhD student (2017-2020), Elise Tourrette M2, doctorante et postdoctorante (2016-2020), Edith Garot Postdoc (2020), Olivier Martin Senior Investigator (2010-2019), Kamel Jebreen postdoctoral fellow (2018-2019), Marianyela Petrizzelli PhD student (2015-2019), Dorian Collot PhD student (2015-2018), Johan Ramsayer (2016), Dephine Sicard (2007-2015), Thelma da Silva PhD student (2011-2014), Timeri Atuhahiva PhD student (2015-2014), Eleonore Durand PhD student (2008-2011), Thibault Nidelet Post-doctoral fellow (2011-2010)Publications
- Raffoux X., Saayman X., Abuelgassim WA., Maret T. , Venon A., Dumas F., Tattini L., Martin OC., Liti G., Falque M. . (2026) Response to divergent selection on meiotic recombination in Saccharomyces cerevisiae. Genetics,
- Ali B., Nicolas S., Blein-Nicolas M. , Martin ML., Djabali Y., Mary-Huard T., Charcosset A., Moreau L., Rincent R.. (2025) Unravelling the Genetic Architecture of Field Traits through Multi-Omics Platform Data integration. Genetics,
- Arenas S., Djabali Y., Rincent R., Cubry P., Martin ML., Blein-Nicolas M. , Laplaze L., Schneider H., Grondin A.. (2025) Modeling plant phenotypic plasticity and its underlying genetic architecture: a comparative study. Journal of Experimental Botany, eraf013
- Blein-Nicolas M. , , La relation génotype-phénotype: des approches réductionnistes aux approches holistiques basées sur la protéomique, HDR, Université Paris Saclay
- Raffoux X., Falque M. . (2025) CAYSS: Package for Automatic Cytometry Analysis of Yeast Spore Segregation. Yeast, 11-12 (41) 681-690
- Andrieu, Marie-Hélène ., Atienza, Isabelle ., Marmagne, Anne ., Just, Daniel ., Bienvenut, Willy Vincent ., Gibon, Yves ., Colombié, Sophie .. (2024) Protocole de marquage à l’azote 15N de fruits de tomate issus de plantes cultivées en condition standard. NOV'AE, 5 (2022) 1-11
- Blein-Nicolas M. , Devijver E., Gallopin M., Perthame E.. (2024) Nonlinear network-based quantitative trait prediction from biological data. Journal of the Royal Statistical Society Series C: Applied Statistics, qlae012
- Falque M. , Bourgais A., Dumas F., De Carvalho M., Diblasi C.. (2024) MiniRead: A simple and inexpensive do‐it‐yourself device for multiple analyses of micro‐organism growth kinetics. Yeast, yea.3932
- Petrizzelli M., Coton C., de Vienne D. . (2024) Formalizing the law of diminishing returns in metabolic networks using an electrical analogy. R. Soc. Open Sci., 10 (11) 240165
- Revillon S., Dillmann C. , Galic N. , Bauland C., Palaffre C., Malvar RA., Butron A., Rebaudo F., Legrand J. . (2024) Effects of maize development and phenology on the field infestation dynamics of the European corn borer (Lepidoptera: Crambidae). Journal of Economic Entomology, 5 (117) 1913-1925
- Urrutia M., Blein-Nicolas M. , Fernandez O., Bernillon S., Maucourt M., Deborde C., Balliau T., Rabier D., Bénard C., Prigent S., Quilleré I., Jacob D., Gibon Y., Zivy M., Giauffret C., Hirel B., Moing A.. (2024) Identification of metabolic and protein markers representative of the impact of mild nitrogen deficit on agronomic performance of maize hybrids. Metabolomics, 6 (20) 128
- Coton C., Dillmann C. , de Vienne D. . (2023) Evolution of enzyme levels in metabolic pathways: A theoretical approach. Part 2. Journal of Theoretical Biology, (558) 111354
- Desbiez-Piat A., Ressayre A. , Marchadier É. , Noly A., Remoué C., Vitte C., Belcram H., Bourgais A., Galic N. , Guilloux ML., Tenaillon MI., Dillmann C. . (2023) Pervasive GxE interactions shape adaptive trajectories and the exploration of the phenotypic space in artificial selection experiments. Genetics, (in press) 2023.01.13.523786
- Djabali Y., Rincent R., Martin ML., Blein-Nicolas M. . (2023) Plasticity QTLs specifically contribute to the genotype × water availability interaction in maize. Theor Appl Genet, 11 (136) 228
- Djabali Yacine ., 13/12/2023, Exploitation des données multi-omiques pour élucider les bases génétiques et moléculaires de caractères complexes : une étude de génétique des systèmes de la réponse du maïs à la sécheresse, PhD, Université Paris Saclay
- Duruflé H., Balliau T., Blanchet N., Chaubet A., Duhnen A., Pouilly N., Blein-Nicolas M. , Mangin B., Maury P., Langlade NB., Zivy M.. (2023) Sunflower Hybrids and Inbred Lines Adopt Different Physiological Strategies and Proteome Responses to Cope with Water Deficit. Biomolecules, 7 (13) 1110
- Michel E., Masson E., Bubbendorf S., Lapicque L., Nidelet T., Segond D., Guézenec S., Marlin T., Devillers H., Rué O., Onno B., Legrand J. , Sicard D.. (2023) Artisanal and farmer bread making practices differently shape fungal species community composition in French sourdoughs. Peer Community Journal, (3) e11
- Plancade S., Marchadier É. , Huet S., Ressayre A. , Noûs C., Dillmann C. . (2023) A successive time-to-event model of phyllochron dynamics for hypothesis testing: application to the analysis of genetic and environmental effects in maize. Plant Methods, 1 (19) 54
- Régnier B., Legrand J. , Calatayud PA., Rebaudo F.. (2023) Developmental Differentiations of Major Maize Stemborers Due to Global Warming in Temperate and Tropical Climates. Insects, 1 (14) 51
- Sanane I., Nicolas SD., Bauland C., Marion-Poll F., Noûs C., Legrand J. , Dillmann C. . (2023) Large genetic variability of maize leaf palatability to european corn borer : metabolic insights. bioRxiv,
- Von Gastrow L., Michel E., Legrand J. , Amelot R., Segond D., Guezenec S., Rué O., Chable V., Goldringer I., Dousset X., Serpolay‐Bessoni E., Taupier‐Letage B., Vindras‐Fouillet C., Onno B., Valence F., Sicard D.. (2023) Microbial community dispersal from wheat grains to sourdoughs: A contribution of participatory research. Molecular Ecology, 10 (32) 2413-2427
- de Vienne D. , Coton C., Dillmann C. . (2023) The genotype–phenotype relationship and evolutionary genetics in the light of the Metabolic Control Analysis. Biosystems, (232) 105000
- Albert B., Matamoro-Vidal A., Prieu C., Nadot S., Till-Bottraud I., Ressayre A. , Gouyon PH.. (2022) A Review of the Developmental Processes and Selective Pressures Shaping Aperture Pattern in Angiosperms. Plants, 3 (11) 357
- Coton C., Talbot G., Le Louarn M., Dillmann C. , de Vienne D. . (2022) Evolution of enzyme levels in metabolic pathways: A theoretical approach. Part 1. Journal of Theoretical Biology, (538) 111015
- David O., Le Rouzic A., Dillmann C. . (2022) Optimization of sampling designs for pedigrees and association studies. Biometrics, 3 (78) 1056-1066
- Hsu YM., 2022-09, Mining genetic diversity for tomorrow's agriculture, PhD, Université Paris-Saclay
- Hsu YM., Falque M. , Martin OC.. (2022) Quantitative modelling of fine-scale variations in the Arabidopsis thaliana crossover landscape. Quant Plant Bio., (3) e3
- Régnier B., Legrand J. , Rebaudo F.. (2022) Modeling Temperature-Dependent Development Rate in Insects and Implications of Experimental Design. Environmental Entomology, 1 (51) 132-144
- de Vienne D. , Capy P.. (2022) Special issue on “The relationship between genotype and phenotype: new insight into an old question”. Genetica, 3-4 (150) 151-151
- de Vienne D. . (2022) What is a phenotype? History and new developments of the concept. Genetica, 3-4 (150) 153-158
- Coton C., 2021-12, Évolution des concentrations d'enzymes dans les réseaux métaboliques, PhD, Université Paris-Saclay
- Desbiez-Piat A., Le Rouzic A., Tenaillon MI., Dillmann C. . (2021) Interplay between extreme drift and selection intensities favors the fixation of beneficial mutations in selfing maize populations. Genetics, 2 (219)
- Desbiez-Piat, 2021, The Dynamics of the Response to Selection under High Drift-High Selection: Insights from Saclay’s Divergent Selection Experiments for Flowering Time in Maize, PhD, Université Paris-Saclay
- Olvera-Vazquez SG., Alhmedi A., Miñarro M., Shykoff JA., Marchadier É. , Rousselet A., Remoué C., Gardet R., Degrave A., Robert P., Chen X., Porchier J., Giraud T., Vander-Mijnsbrugee K., Raffoux X., Falque M. , Alins G., Didelot F., Beliën T., Dapena E., Lemarquand A., Cornille A.. (2021) Experimental test for local adaptation of the rosy apple aphid (Dysaphis plantaginea) to its host (Malus domestica) and to its climate in Europe. PCI Ecology, (Pre-registration version)
- Petrizzelli MS., de Vienne D. , Nidelet T., Noûs C., Dillmann C. . (2021) Data integration uncovers the metabolic bases of phenotypic variation in yeast. PLoS Comput Biol, 7 (17) e1009157
- Plancade S., Marchadier É. , Huet S., Ressayre A. , Noûs C., Dillmann C. . (2021) A new hypothesis-testing model for phyllochron based on a stochastic process - application to analysis of genetic and environment effects in maize. DOI.org (Crossref),
- Sanané I., Legrand J. , Dillmann C. , Marion-Poll F.. (2021) High-Throughput Feeding Bioassay for Lepidoptera Larvae. J Chem Ecol, 7 (47) 642-652
- Tourrette E., Falque M. , Martin OC.. (2021) Enhancing backcross programs through increased recombination. Genet Sel Evol, 1 (53) 25
- Blein-Nicolas M. , Negro SS., Balliau T., Welcker C., Cabrera-Bosquet L., Nicolas SD., Charcosset A., Zivy M.. (2020) A systems genetics approach reveals environment-dependent associations between SNPs, protein coexpression, and drought-related traits in maize. Genome Res., 11 (30) 1593-1604
- Carrillo-Perdomo E., Vidal A., Kreplak J., Duborjal H., Leveugle M., Duarte J., Desmetz C., Deulvot C., Raffiot B., Marget P., Tayeh N., Pichon JP., Falque M. , Martin OC., Burstin J., Aubert G.. (2020) Development of new genetic resources for faba bean (Vicia faba L.) breeding through the discovery of gene-based SNP markers and the construction of a high-density consensus map. Sci Rep, 1 (10) 6790
- Falque M. , Jebreen K., Paux E., Knaak C., Mezmouk S., Martin OC.. (2020) CNVmap: A Method and Software To Detect and Map Copy Number Variants from Segregation Data. Genetics, 3 (214) 561-576
- Goldringer I., van Frank G., Bouvier d’Yvoire C., Forst E., Galic N. , Garnault M., Locqueville J., Pin S., Bailly J., Baltassat R., Berthellot JF., Caizergues F., Dalmasso C., de Kochko P., Gascuel JS., Hyacinthe A., Lacanette J., Mercier F., Montaz H., Ronot B., Rivière P.. (2020) Agronomic Evaluation of Bread Wheat Varieties from Participatory Breeding: A Combination of Performance and Robustness. Sustainability, 1 (12) 128
- Hanemian M., Vasseur F., Marchadier É. , Gilbault E., Bresson J., Gy I., Violle C., Loudet O.. (2020) Natural variation at FLM splicing has pleiotropic effects modulating ecological strategies in Arabidopsis thaliana. Nat Commun, 1 (11) 4140
- Harlé O., Legrand J. , Tesnière C., Pradal M., Mouret JR., Nidelet T.. (2020) Investigations of the mechanisms of interactions between four non-conventional species with Saccharomyces cerevisiae in oenological conditions. PLoS ONE, 5 (15) e0233285
- Henry V., Saïs F., Inizan O., Marchadier É. , Dibie J., Goelzer A., Fromion V.. (2020) BiPOm: a rule-based ontology to represent and infer molecule knowledge from a biological process-centered viewpoint. BMC Bioinformatics, 1 (21) 327
- Sanane I., 20 octobre 2020, Composantes de la dynamique de l’interaction entre le maïs et les insectes lépidoptères foreurs de tige, PhD, Université Paris-Saclay
- de Vienne D. , Fiévet JB.. (2020) The Pitfalls of Heterosis Coefficients. Plants, 7 (9) 875
- Bienvenut W. V. , 2 décembre 2019, Des débuts de la protéomique à l'acétylation N-terminale des protéines chez les plantes, HDR, Université Paris Saclay
- Jebreen K., Petrizzelli M., Martin OC.. (2019) Probabilities of Multilocus Genotypes in SIB Recombinant Inbred Lines. Front. Genet., (10) 833
- Petrizzelli M., de Vienne D. , Dillmann C. . (2019) Decoupling the Variances of Heterosis and Inbreeding Effects Is Evidenced in Yeast’s Life-History and Proteomic Traits. Genetics, 2 (211) 741-756
- Petrizzelli M., 2019-07, Mathematical modelling and integration of complex biological data : analysis of the heterosis phenomenon in yeast, PhD, Université Paris Saclay (COmUE)
- Srisuwan S., Sihachakr D., Martin J., Valles J., Ressayre A. , Brown SC., Siljak-Yakovlev S.. (2019) Change in nuclear DNA content and pollen size with polyploidisation in the sweet potato (Ipomoea batatas, Convolvulaceae) complex. Plant Biol (Stuttg), 2 (21) 237-247
- Tenaillon MI., Seddiki K., Mollion M., Guilloux ML., Marchadier É. , Ressayre A. , Dillmann C. . (2019) Transcriptomic response to divergent selection for flowering times reveals convergence and key players of the underlying gene regulatory network. bioRxiv, 461947
- Termolino P., Falque M. , Aiese Cigliano R., Cremona G., Paparo R., Ederveen A., Martin OC., Consiglio FM., Conicella C.. (2019) Recombination suppression in heterozygotes for a pericentric inversion induces the interchromosomal effect on crossovers in Arabidopsis. Plant J, 6 (100) 1163-1175
- Tourrette E., Bernardo R., Falque M. , Martin OC.. (2019) Assessing by Modeling the Consequences of Increased Recombination in Recurrent Selection of Oryza sativa and Brassica rapa. G3, 12 (9) 4169-4181
- Tourrette E., 25 novembre 2019, Unleashing genetic diversity in breeding by increasing recombination: an in silico study, PhD, Université de Paris-Diderot
- Urien C., Legrand J. , Montalent P., Casaregola S., Sicard D.. (2019) Fungal Species Diversity in French Bread Sourdoughs Made of Organic Wheat Flour. Front. Microbiol., (10) 201
- Vasseur F., Fouqueau L., de Vienne D. , Nidelet T., Violle C., Weigel D.. (2019) Nonlinear phenotypic variation uncovers the emergence of heterosis in Arabidopsis thaliana. PLoS Biol, 4 (17) e3000214
- Albert B., Ressayre A. , Dillmann C. , Carlson AL., Swanson RJ., Gouyon PH., Dobritsa AA.. (2018) Effect of aperture number on pollen germination, survival and reproductive success in Arabidopsis thaliana. Ann. Bot., 4 (121) 733-740
- Carbonetto B., Ramsayer J., Nidelet T., Legrand J. , Sicard D.. (2018) Bakery yeasts, a new model for studies in ecology and evolution. Yeast, 11 (35) 591-603
- Collot D., Nidelet T., Ramsayer J., Martin OC., Meleard S., Dillmann C. , Sicard D., Legrand J. . (2018) Feedback between environment and traits under selection in a seasonal environment: consequences for experimental evolution. Proceedings. Biological sciences, 1876 (285)
- Collot D., 2018-06-19 19/06/18, Modélisation des dynamiques adaptatives de la levure de boulanger S. cerevisae dans un environnement saisonnier, PhD, Université Paris-Saclay
- Fiévet JB., Nidelet T., Dillmann C. , de Vienne D. . (2018) Heterosis Is a Systemic Property Emerging From Non-linear Genotype-Phenotype Relationships: Evidence From in Vitro Genetics and Computer Simulations. Front. Genet., (9)
- Grigolon S., Bravi B., Martin OC.. (2018) Responses to auxin signals: an operating principle for dynamical sensitivity yet high resilience. Royal Society Open Science, 1 (5) 172098
- Raffoux X., 2018-11, Diversité et déterminisme génétique de la recombinaison méiotique chez Saccharomyces cerevisiae, PhD, Université Paris Saclay (COmUE)
- Raffoux X., Bourge M., Dumas F., Martin OC., Falque M. . (2018) High-throughput measurement of recombination rates and genetic interference in Saccharomyces cerevisiae. Yeast, 6 (35) 431-442
- Raffoux X., Bourge M., Dumas F., Martin OC., Falque M. . (2018) Role of Cis, Trans, and Inbreeding Effects on Meiotic Recombination in Saccharomyces cerevisiae. Genetics, 4 (210) 1213-1226
- Pelé A., Falque M. , Trotoux G., Eber F., Nègre S., Gilet M., Huteau V., Lodé M., Jousseaume T., Dechaumet S., Morice J., Poncet C., Coriton O., Martin OC., Rousseau-Gueutin M., Chèvre AM.. (2017) Amplifying recombination genome-wide and reshaping crossover landscapes in Brassicas. PLoS Genet, 5 (13)
- Samal A., Martin OC.. (2017) Haldane, Waddington and recombinant inbred lines: extension of their work to any number of genes. J. Genet., 5 (96) 795-800
- Henry A., Martin OC.. (2016) Short relaxation times but long transient times in both simple and complex reaction networks. Journal of the Royal Society, Interface / the Royal Society, 120 (13)
- Hosseini SR., Martin OC., Wagner A.. (2016) Phenotypic innovation through recombination in genome-scale metabolic networks. Proceedings. Biological sciences, 1839 (283)
- Jacques N., Sarilar V., Urien C., Lopes MR., Morais CG., Uetanabaro AP., Tinsley CR., Rosa CA., Sicard D., Casaregola S.. (2016) Three novel ascomycetous yeast species of the Kazachstania clade, Kazachstania saulgeensis sp. nov., Kazachstania serrabonitensis sp. nov. and Kazachstania australis sp. nov. Reassignment of Candida humilis to Kazachstania humilis f.a. comb. nov. and Candida pseudohumilis to Kazachstania pseudohumilis f.a. comb. nov. International journal of systematic and evolutionary microbiology, 12 (66) 5192-5200
- Legrand J. , Bolotin-Fukuhara M., Bourgais A., Fairhead C. , Sicard D.. (2016) Life-history strategies and carbon metabolism gene dosage in the Nakaseomyces yeasts. FEMS yeast research, 2 (16) fov112
- Lhomme E., Urien C., Legrand J. , Dousset X., Onno B., Sicard D.. (2016) Sourdough microbial community dynamics: An analysis during French organic bread-making processes. Food Microbiology, Pt A (53) 41-50
- Martin OC., Krzywicki A., Zagorski M.. (2016) Drivers of structural features in gene regulatory networks: From biophysical constraints to biological function. Physics of life reviews, (17) 124-58
- Mori M., Hwa T., Martin OC., De Martino A., Marinari E.. (2016) Constrained Allocation Flux Balance Analysis. PLoS computational biology, 6 (12) e1004913
- Sabarly V., Aubron C., Glodt J., Balliau T., Langella O., Chevret D., Rigal O., Bourgais A., Picard B., de Vienne D. , Denamur E., Bouvet O., Dillmann C. . (2016) Interactions between genotype and environment drive the metabolic phenotype within Escherichia coli isolates. Environmental microbiology, 1 (18) 100-17
- Blein-Nicolas M. , Albertin W., da Silva T., Valot B., Balliau T., Masneuf-Pomarède I., Bely M., Marullo P., Sicard D., Dillmann C. , de Vienne D. , Zivy M.. (2015) A Systems Approach to Elucidate Heterosis of Protein Abundances in Yeast. Mol. Cell Proteomics, 8 (14) 2056-2071
- Durand E., Tenaillon MI., Raffoux X., Thépot S., Falque M. , Jamin P., Bourgais A., Ressayre A. , Dillmann C. . (2015) Dearth of polymorphism associated with a sustained response to selection for flowering time in maize. BMC Evol Biol, 1 (15) 103
- Falque M. , 2015-10-15 15/10/15, Sur les traces de la recombinaison méiotique, source de biodiversité et outil génétique, HDR, Université Paris-Sud
- Grigolon S., Sollich P., Martin OC.. (2015) Modelling the emergence of polarity patterns for the intercellular transport of auxin in plants. Journal of the Royal Society, Interface / the Royal Society, 106 (12)
- Henry A., 2015-12-17 17/12/15, Modélisation cinétique du métabolisme : construction du modèle, analyse et applications biotechnologiques, PhD, Université Paris Diderot - Paris 7
- Sidhu GK., Fang C., Olson MA., Falque M. , Martin OC., Pawlowski WP.. (2015) Recombination patterns in maize reveal limits to crossover homeostasis. PNAS, 52 (112) 15982-15987
- Urien C., 2015-01, Diversité des espèces de levures dans des levains naturels français produits à partir de farine issue de l'Agriculture Biologique : une étude pilote pour analyser les pratiques boulangères et les patterns des communautés microbiennes, PhD, Université Paris Sud - Paris XI
- da Silva T., Albertin W., Dillmann C. , Bely M., la Guerche S., Giraud C., Huet S., Sicard D., Masneuf-Pomarede I., de Vienne D. , Marullo P.. (2015) Hybridization within Saccharomyces Genus Results in Homoeostasis and Phenotypic Novelty in Winemaking Conditions. PloS one, 5 (10) e0123834
- Anderson LK., Lohmiller LD., Tang X., Hammond DB., Javernick L., Shearer L., Basu-Roy S., Martin OC., Falque M. . (2014) Combined fluorescent and electron microscopic imaging unveils the specific properties of two classes of meiotic crossovers. Proceedings of the National Academy of Sciences of the United States of America, 37 (111) 13415-20
- Jahns MT., Vezon D., Chambon A., Pereira L., Falque M. , Martin OC., Chelysheva L., Grelon M.. (2014) Crossover localisation is regulated by the neddylation posttranslational regulatory pathway. PLoS biology, 8 (12) e1001930
- Roy S., 2014-03-31 31/03/14, Models of meiotic crossover formation, heterogeneity and inter-pathway cross-talks, PhD thesis, Université Paris Sud
- Spor A., Kvitek DJ., Nidelet T., Martin J., Legrand J. , Dillmann C. , Bourgais A., de Vienne D. , Sherlock G., Sicard D.. (2014) Phenotypic and genotypic convergences are influenced by historical contingency and environment in yeast. Evolution; international journal of organic evolution, 3 (68) 772-90
- Suay L., Zhang DS., Eber F., Jouy H., Lode M., Huteau V., Coriton O., Szadkowski E., Leflon M., Martin OC., Falque M. , Jenczewski E., Paillard S., Chevre AM.. (2014) Crossover rate between homologous chromosomes and interference are regulated by the addition of specific unpaired chromosomes in Brassica. The New phytologist, 2 (201) 645-656
- Albertin W., Marullo P., Bely M., Aigle M., Bourgais A., Langella O., Balliau T., Chevret D., Valot B., da Silva T., Dillmann C. , de Vienne D. , Sicard D.. (2013) Linking post-translational modifications and variation of phenotypic traits. Molecular & cellular proteomics : MCP, 3 (12) 720-35
- Albertin W., da Silva T., Rigoulet M., Salin B., Masneuf-Pomarede I., de Vienne D. , Sicard D., Bely M., Marullo P.. (2013) The mitochondrial genome impacts respiration but not fermentation in interspecific Saccharomyces hybrids. PloS one, 9 (8) e75121
- Basu-Roy S., Gauthier F., Giraut L., Mezard C., Falque M. , Martin OC.. (2013) Hot Regions of Noninterfering Crossovers Coexist with a Nonuniformly Interfering Pathway in Arabidopsis thaliana. Genetics, 3 (195) 769-79
- Bauer E., Falque M. , Walter H., Bauland C., Camisan C., Campo L., Meyer N., Ranc N., Rincent R., Schipprack W., Altmann T., Flament P., Melchinger AE., Menz M., Moreno-Gonzalez J., Ouzunova M., Revilla P., Charcosset A., Martin OC., Schon CC.. (2013) Intraspecific variation of recombination rate in maize. Genome biology, 9 (14) R103
- Blein-Nicolas M. , Albertin W., Valot B., Marullo P., Sicard D., Giraud C., Huet S., Bourgais A., Dillmann C. , de Vienne D. , Zivy M.. (2013) Yeast proteome variations reveal different adaptive responses to grape must fermentation. Molecular biology and evolution, 6 (30) 1368-83
- Drouaud J., Khademian H., Giraut L., Zanni V., Bellalou S., Henderson IR., Falque M. , Mezard C.. (2013) Contrasted Patterns of Crossover and Non-crossover at Arabidopsis thaliana Meiotic Recombination Hotspots. PLoS genetics, 11 (9) e1003922
- Blein-Nicolas M. , Xu H., de Vienne D. , Giraud C., Huet S., Zivy M.. (2012) Including shared peptides for estimating protein abundances: a significant improvement for quantitative proteomics. Proteomics, 18 (12) 2797-801
- Durand E., Bouchet S., Bertin P., Ressayre A. , Jamin P., Charcosset A., Dillmann C. , Tenaillon MI.. (2012) Flowering time in maize: linkage and epistasis at a major effect locus. Genetics, 4 (190) 1547-62
- Albertin W., Marullo P., Aigle M., Dillmann C. , de Vienne D. , Bely M., Sicard D.. (2011) Population size drives industrial Saccharomyces cerevisiae alcoholic fermentation and is under genetic control. Applied and environmental microbiology, 8 (77) 2772-84
- Fiévet J., de Vienne D. , Gallais A.. (2011) Heterosis, Inbreeding depression and Hybrid varieties. library.siena.edu,
- Gauthier F., Martin O., Falque M. . (2011) CODA (crossover distribution analyzer): quantitative characterization of crossover position patterns along chromosomes. BMC Bioinformatics, 1 (12) 27
- Giraut L., Falque M. , Drouaud J., Pereira L., Martin OC., Mezard C.. (2011) Genome-wide crossover distribution in Arabidopsis thaliana meiosis reveals sex-specific patterns along chromosomes. PLoS genetics, 11 (7) e1002354
- Sabarly V., Bouvet O., Glodt J., Clermont O., Skurnik D., Diancourt L., de Vienne D. , Denamur E., Dillmann C. . (2011) The decoupling between genetic structure and metabolic phenotypes in Escherichia coli leads to continuous phenotypic diversity. Journal of evolutionary biology, 7 (24) 1559-71
- Wang S., Spor A., Nidelet T., Montalent P., Dillmann C. , de Vienne D. , Sicard D.. (2011) Switch between life history strategies due to changes in glycolytic enzyme gene dosage in Saccharomyces cerevisiae. Applied and environmental microbiology, 2 (77) 452-9



















